RFP Lab
purpose- Make RFP from jelly fish in bacteria. To also learn about steps in genetic engineering
Materials + Procedure: These can be found in their corresponding areas in the Amgen lab manual.(Ex. 2a materials and procedure are in part 2a for the RFP lab)
Materials + Procedure: These can be found in their corresponding areas in the Amgen lab manual.(Ex. 2a materials and procedure are in part 2a for the RFP lab)
OVERVIEW
2a: Preparing the RFP Gene
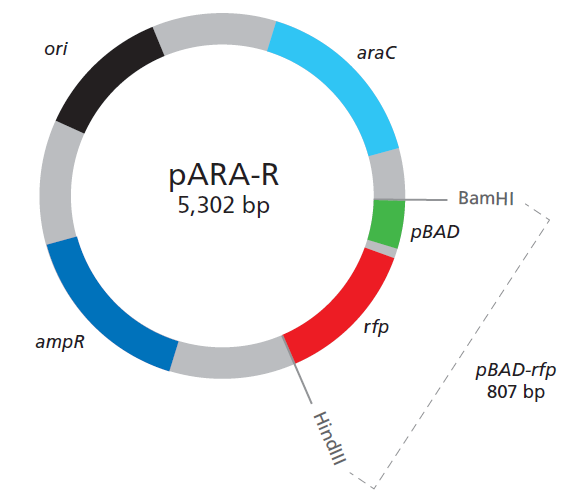
- In this first lab we separated the RFP-Ara gene from the pAra-R plasmid using BamH1 and HindIII
4a: Electrophoresis to verify plasmid
- Electrophoresis was used to make sure that we had cut the plasmid correctly. We ran the gene and plasmid along with a DNA ladder that will tell you the molecular sizes they should be. We then used the ladder to compare our cut sizes and what they should be.
5a: Transformation of Bacteria using a Recombinant Plasmid
- we took restriction enzymes to cut the plasmid and ligase to paste the RFP gene. We then put on a selective marker to regulate the growth of bacteria that were not resistant to things like Ampicillin. The bacteria were transformed into a recombinant plasmid.
6a: Separating the RFP Gene with Chromatography
- We used a chromatography column to separate the RFP gene from everything else. There were beads in the column that we used to catch the RFP to seperate it and then extract the gene from the beads with a buffer.
2a: Preparing the RFP Gene
- In this first lab we separated the RFP-Ara gene from the pAra-R plasmid using BamH1 and HindIII
4a: Electrophoresis to verify plasmid
- Electrophoresis was used to make sure that we had cut the plasmid correctly. We ran the gene and plasmid along with a DNA ladder that will tell you the molecular sizes they should be. We then used the ladder to compare our cut sizes and what they should be.
5a: Transformation of Bacteria using a Recombinant Plasmid
- we took restriction enzymes to cut the plasmid and ligase to paste the RFP gene. We then put on a selective marker to regulate the growth of bacteria that were not resistant to things like Ampicillin. The bacteria were transformed into a recombinant plasmid.
6a: Separating the RFP Gene with Chromatography
- We used a chromatography column to separate the RFP gene from everything else. There were beads in the column that we used to catch the RFP to seperate it and then extract the gene from the beads with a buffer.
Results
Lab 2A
- We cut a plasmid, extracting the RFP-Ara gene, and ran it through a gel. There, we saw that the size of the molecule was accurate -- it corresponded with the DNA ladder.
Lab 4a
If you don't have the correct recombinant plasmid the damaged bacteria will go unnoticed because it might not be visible, therefore you would have to restart the lab.
My gel results were the same as my predictions because they had the same number of bands in each lane as I had predicted.
There is a couple of bands unexpected and this could be a result of either human error or we damaged the gel with the DNA.
Yes because we got the correct results and the R+ lanes had two fragments, which was desired.
Yes, I saw multiple different bands showing that the plasmid was digested.
Yes, there were multiple bands showing the plasmid was digested into multiple fragments.
The RFP gene would be found in the R+ lane and the ampR gene in R- lane.We can locate these in both of these lanes.
The lanes with plasmids are supercoiled and both migrate. Yes the fragments have elongated edges while the plasmids are supercoiled.
- We cut a plasmid, extracting the RFP-Ara gene, and ran it through a gel. There, we saw that the size of the molecule was accurate -- it corresponded with the DNA ladder.
Lab 4a
If you don't have the correct recombinant plasmid the damaged bacteria will go unnoticed because it might not be visible, therefore you would have to restart the lab.
My gel results were the same as my predictions because they had the same number of bands in each lane as I had predicted.
There is a couple of bands unexpected and this could be a result of either human error or we damaged the gel with the DNA.
Yes because we got the correct results and the R+ lanes had two fragments, which was desired.
Yes, I saw multiple different bands showing that the plasmid was digested.
Yes, there were multiple bands showing the plasmid was digested into multiple fragments.
The RFP gene would be found in the R+ lane and the ampR gene in R- lane.We can locate these in both of these lanes.
The lanes with plasmids are supercoiled and both migrate. Yes the fragments have elongated edges while the plasmids are supercoiled.
Questions: 5A
1. The P+ bacteria culture is treated differently because it has more plates than P- for the sake of comparing results. The P- culture is there to see the effects of LB and amp to the bacteria.
2. Cells need time to recover after the heat shock, otherwise they might die if they can't re-stabilize.
3. Cells are incubated at 37 degrees C because that is a normal temperature condition.
4. Aseptic is important in this lab because it keeps the bacteria from contaminating and becoming contaminated itself.
5. Most of my results except for the P+ only plate matched my predictions. I realized that the P+ was resistant to ara and amp, unlike I thought they were.
6. There were no red colonies on the LB/amp/ara plate.
7. The red colonies might only appear on the LB/amp/ara plate and not the LB/amp plate because it needs ara to survive and make the rfp.
8. It is important to have many copies of a recombinant plasmid in a cell to create more protein in a cell.
9. The rfp gene makes specialized proteins that develop into the traits of an organism.
10. Bacteria can make any protein because they are made to produce proteins, and if they are given the right codes, they will make any proteins.
1. The P+ bacteria culture is treated differently because it has more plates than P- for the sake of comparing results. The P- culture is there to see the effects of LB and amp to the bacteria.
2. Cells need time to recover after the heat shock, otherwise they might die if they can't re-stabilize.
3. Cells are incubated at 37 degrees C because that is a normal temperature condition.
4. Aseptic is important in this lab because it keeps the bacteria from contaminating and becoming contaminated itself.
5. Most of my results except for the P+ only plate matched my predictions. I realized that the P+ was resistant to ara and amp, unlike I thought they were.
6. There were no red colonies on the LB/amp/ara plate.
7. The red colonies might only appear on the LB/amp/ara plate and not the LB/amp plate because it needs ara to survive and make the rfp.
8. It is important to have many copies of a recombinant plasmid in a cell to create more protein in a cell.
9. The rfp gene makes specialized proteins that develop into the traits of an organism.
10. Bacteria can make any protein because they are made to produce proteins, and if they are given the right codes, they will make any proteins.
Lab 6a
In binding spaces proteins attach to other molecules and this allows it to carry out its function.
The sequence of the amino acids relates to folding.
The Elute is brighter than the lysate. The RFP will account for that.
Hydrophobic amino acids are used as the basis for separation by column chromatography
Use more of the original buffer to increase the purity of the red fluorescent protein sample.
In binding spaces proteins attach to other molecules and this allows it to carry out its function.
The sequence of the amino acids relates to folding.
The Elute is brighter than the lysate. The RFP will account for that.
Hydrophobic amino acids are used as the basis for separation by column chromatography
Use more of the original buffer to increase the purity of the red fluorescent protein sample.
Analasys Our band was The fifth from the right. The purity of the gene was not very pure, and the thickest band is about where it should have been, but it was not as dark as it should have been. These results could have been due to an error in the process of isolation the RFP, or in the process of multiplying it.